H5N1 model helps predicting the spread of H5N8
Published on February 02, 2017, by Marius Gilbert, Jean Artois, Madhur Dhingra & Catherine Linard
(Updated 2nd Feb. 2017) We evaluated the predictive capacity of our global H5N1 suitability model published a few month ago in e-life, and based on HPAI H5N1 and H5Nx records of years 2006-2015 in its capacity to predict the current wave of HPAI H5N8 across Europe.
On February 1st, we extracted all the winter 2016:2017 H5N8 HPAI cases in domestic poultry (i.e. excluding wild bird cases) from the Empres-I database. In the figure below, the cases are distributed over the HPAI H5N1 model (left) and HPAI H5Nx clade 2.3.4.4 model (right).
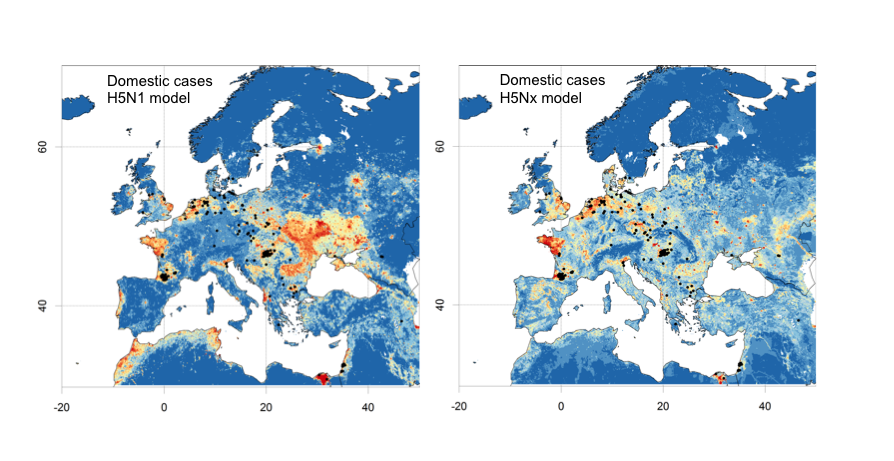
In order to quantify the goodness of fit, we estimated the area under curve (AUC) in two separate ways. First by considering all pseudo-absences in Europe, and second by considering only pseudo-absences located within 150 km of any HPAI H5N8 wild bird case, as illustrated in the figure and tables below.
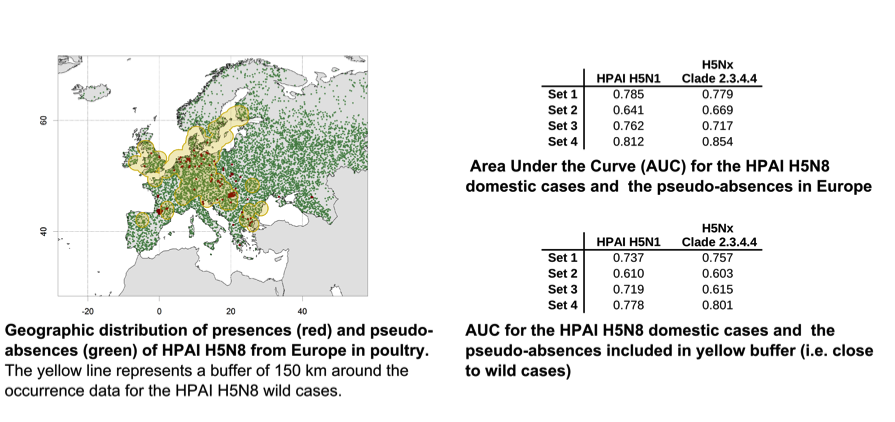

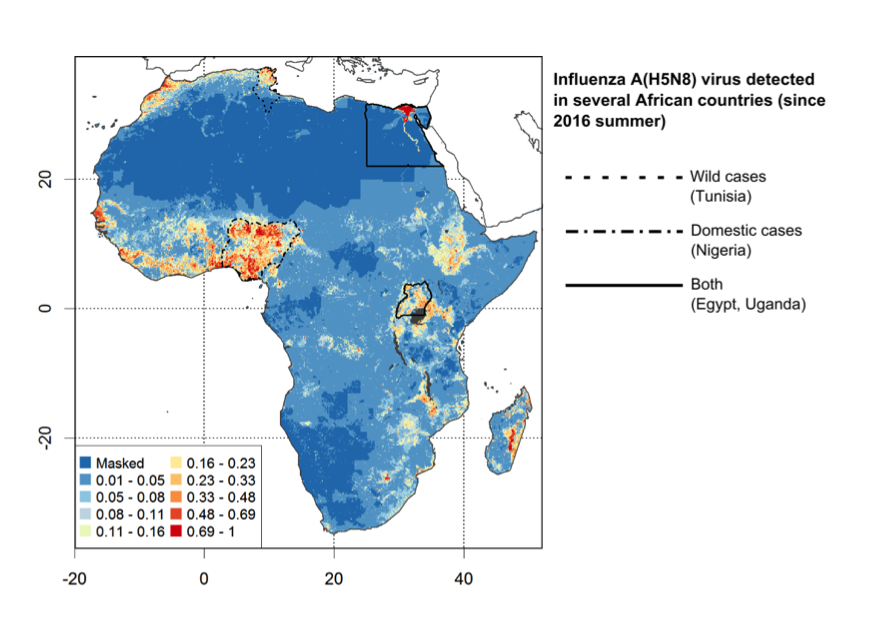
The results highlight that the HPAI H5Nx clade 2.3.4.4 model predictions of predictor Set 4 (a combination of host-related and environmental variables) had the best fit, The AUC when considering all pseudo-absences (all green dots) or only pseudo-absences within 150 km of any wild bird (green dots within the yellow buffer) cases were 0.854 and 0.801, respectively. Since the H5N1 model was found to have the highest extrapolation capacity in our paper, the figure below shows the suitabilty area predicted by the HPAI H5N1 model in sub-saharan Africa, where it may inform surveillance and early detection.
Reference
Global mapping of highly pathogenic avian influenza H5N1 and H5Nx clade 2.3.4.4 viruses with spatial cross-validation M.S. Dhingra, J. Artois, T.P. Robinson, C. Linard, C. Chaiban, I. Xenarios, R. Engler, R. Liechti, D. Kuznetsov, X. Xiao, S.V. Dobschuetz, F. Claes, S.H. Newman, G. Dauphin and M. Gilbert. In “eLife”, vol. 5, 2016