New publication in Science: spatio-temporal invasion dynamics of SARS-CoV-2 Alpha variant in UK
Published on July 22, 2021, by Simon Dellicour
Understanding the causes and consequences of the emergence of SARS-CoV-2 variants of concern is crucial to pandemic control yet difficult to achieve, as they arise in the context of variable human behaviour and immunity. We investigate the spatial invasion dynamics of lineage B.1.1.7 by jointly analysing UK human mobility, virus genomes, and community-based PCR data. We identify a multi-stage spatial invasion process in which early B.1.1.7 growth rates were associated with mobility and asymmetric lineage export from a dominant source location, enhancing the effects of B.1.1.7’s increased intrinsic transmissibility. We further explore how B.1.1.7 spread was shaped by non-pharmaceutical interventions and spatial variation in previous attack rates. Our findings show that careful accounting of the behavioural and epidemiological context within which variants of concern emerge is necessary to interpret correctly their observed relative growth rates. Read the whole study here.
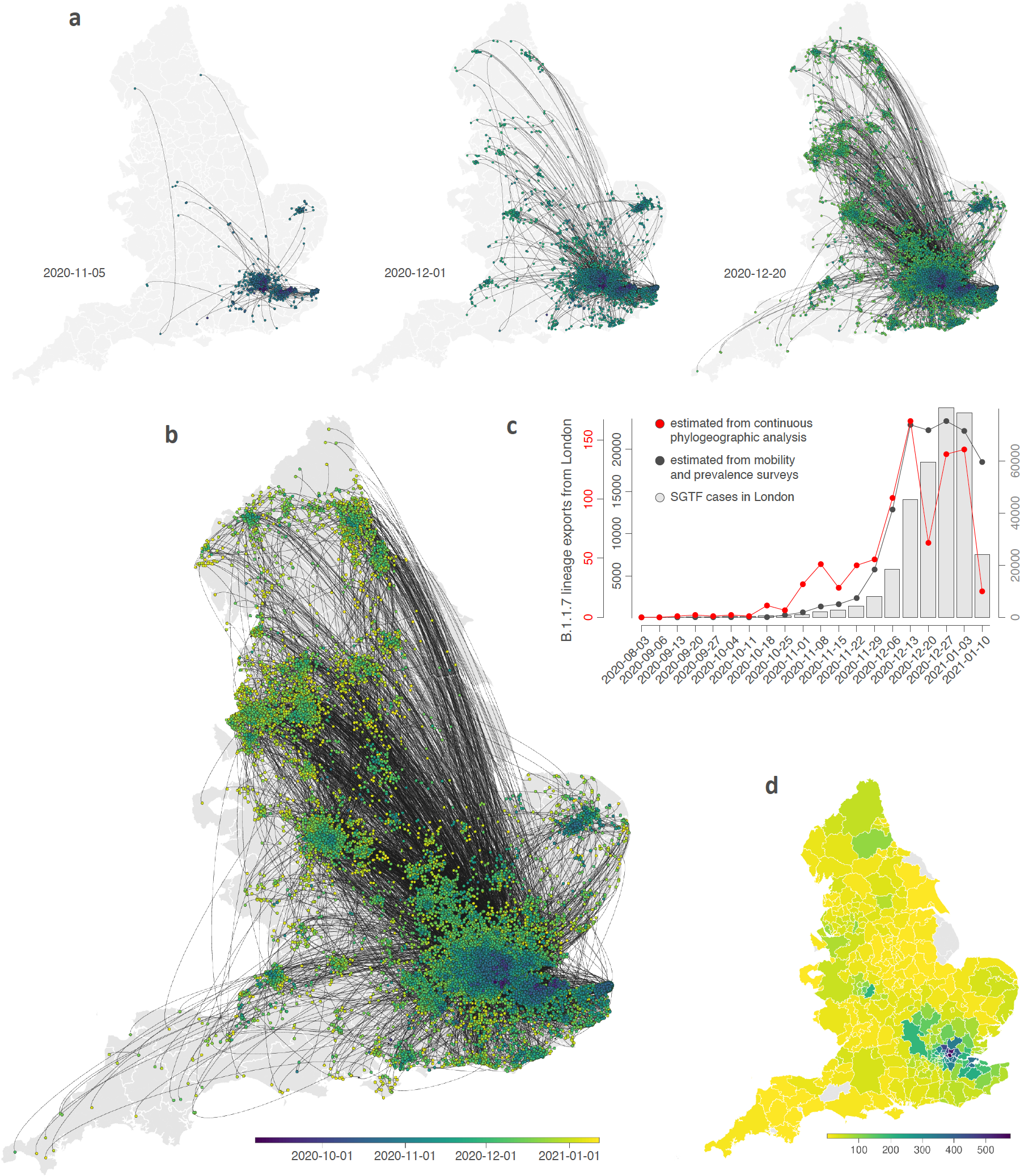
Reference: Kraemer MUG*, Hill V*, Ruis C*, Dellicour S*, Bajaj S*, McCrone JT, Baele G, Parag KV, Lindström Battle A, Gutierrez B, Jackson B, Colquhoun R, O’Toole Á, Klein B, Vespignani A, The COVID-19 Genomics UK (CoG-UK) Consortium, Volz E, Faria NF, Aanensen D, Loman NJ, du Plessis L, Cauchemez S, Rambaut A, Scarpino SV, Pybus OG (2021). Spatio-temporal invasion dynamics of SARS-CoV-2 lineage B.1.1.7 emergence. Science 373: 889-895