Livestock diseases and mapping
Studying livestock diseases
The largest body of our research has dealt with avian influenza (“bird flu”) H5N1 and H7N9 in Asia. We have also been studying the spatial epidemiology of several other livestock diseases at both local and continental scales to better understand their spatial distribution and to identify their main spatial drivers.
In 2004, the highly pathogenic avian influenza (HPAI) H5N1 virus started spreading throughout Asia, causing the death of millions of poultry and transmitting occasionaly to humans, with some fatalities. In the following years, the virus spread across Eurasia down to Africa. Today, it persists in a limited set of countries (Egypt, Bangladesh, China, Vietnam, Indonesia). Our work consisted in identifying the main risk factors associated with the presence or absence of the diseases at multiple scale, in particular the role of host species, and type of farming. Recently, a different avian influenza H7N9 virus emerged in China, low pathogenic in poultry this time, but causing human fatalities too.
Our current research aims to understand the agro-ecological drivers of avian influenza emergence. Why are these viruses emerging ? Where are they more likely to emerge ? Why are they showing a higher apparent seasonality ? What is the respective role of wild bird connectivity and trade in pattern of virus migrations ? These are the numerous questions we are trying to tackle through geospatial analysis and modelling.
Our work also entails producting suitability maps at different spatial scale that can be used to be target suveillance and control in high-risk areas. In the long run, we aim to better understand the role of changes in production and trade systems in the emergence and spread of those viruses, so that we can project how changes in those systems resulting from intensification of the production may translate into disease risk. In addition, we aim to better complement geospatial approaches with phylogeographic analysis and inference to better understand how different factors may shape the evolution and spread of these viruses.
Previous work has involved studies on bovine tuberculosis in Belgium and in the UK, and on the vectors of Bluetongue disease in Belgium and Italy. We currently work on applying spatial modelling to different disease such as Bluetongue in Europe, Porcine Reproductive and Respiratory Syndrome (PRRS), Foot and Mouth Disease (FMD) and Nipah virus infections in Thailand.

Mapping livestock
For many years, several research groups and international organisations have developed high-resolution global maps of livestock based on reported statistics derived from census and survey data. Due to the dynamic nature of these populations (intensification of livestock production) this is an extremely challenging task that involves the collection of contemporary statistics, the development of complex algorithms to disaggregate these statistics at higher spatial resolutions and make predictions where they are absent.
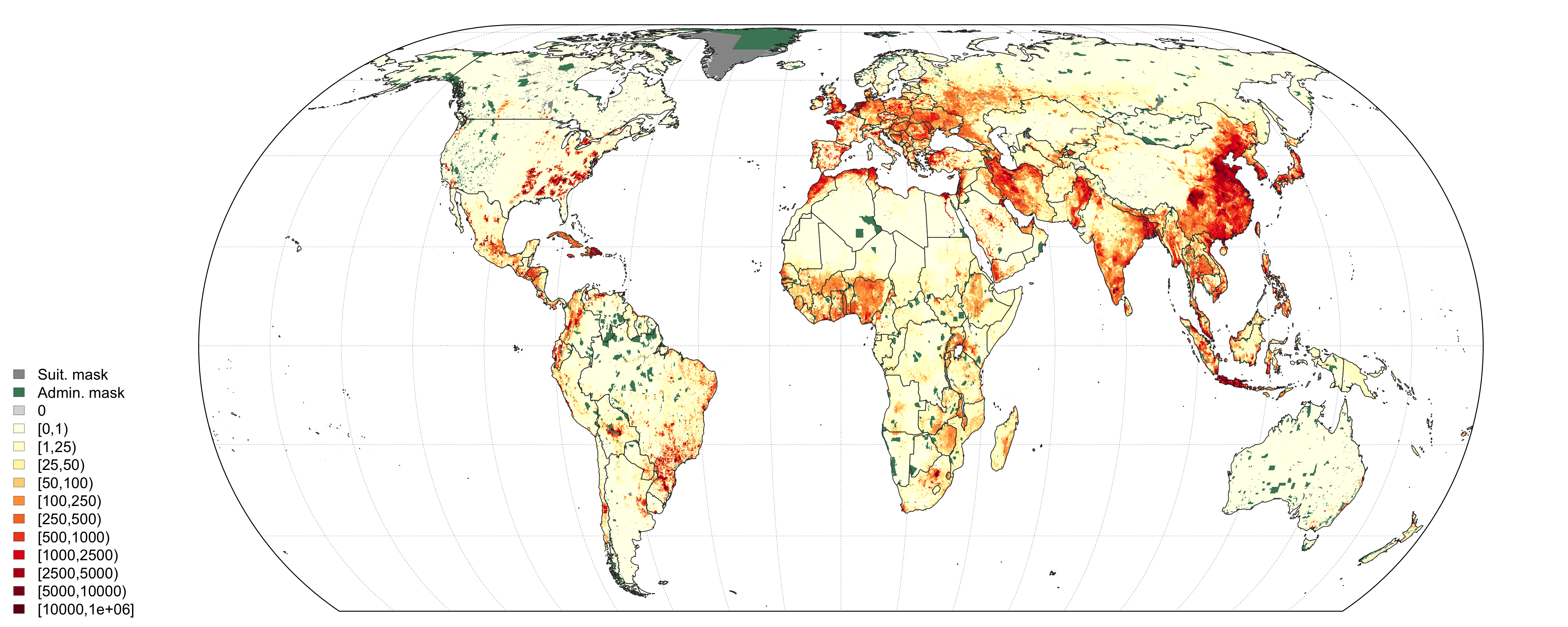
Global output distribution of chicken density of the Gridded Livestock of the World (GLW3) (Gilbert et al. 2018)
The main objectives of our ongoing work are to evaluate quantitatively and qualitatively the different approaches that are used to predict and represent livestock distribution in high-resolution global databases. Our initial efforts focused on the establishment of global livestock distribution data representing the best data available to date (Gilbert et al. 2018).
We now work on the development of time series for the past (1990 - present) and for the future (present - 2050) based on time series of predictor variables and on adapted modelling procedures that can take advantages of training data in space and time.
In parrallel, we also worked on the development of different models that are better suited to predict monogastric species (poultry and pigs) that can be raised in very high number within a single locations. We investigated the use of point-pattern simulation modelling to predict farm locations (Chaiban et al. 2019) and ocne calibrated, the approach will be applied to create realistic synthetic farm distribution models in relevant countries.
Further developments involve predicting how intensification of the livestock production sector may impact the geoghraphical distribution of animals and farms in ruminant and monogastric livestock production systems. We also work toward the development of animal movement models (Nicolas et al. 2018) that can be used to predict movement networks.
Our work is carried out in collaboration with the FAO Livestock and Environment group, and is presented in FAO Livestock Systems web page. The data themselves are distributed through the Global Livestock Data Harvard dataverse.
Collaborations
Our main institutional collaborators are or have been T. Robinson (ILRI, Nairobi, Kenya), G. Dauphin, W. Kalpravidth, S. Newman, V. Martin, J. Slingenbergh, G. Cinardi (FAO, Rome, Italy), W. Thanapongtharm (Department of Livestock Development, DLD, Bangkok, Thailand), H. Yu (Chinese Center for Disease Control, CDC, Beijing, China), A. Conte & Carla Ippoliti (IZS Teramo), and our main academic collaborators are or have been Xiangming Xiao (Univ. Oklahoma, USA), Julien Cappelle (CIRAD, Montpellier, France), D. Pfeiffer (Royal Veterinary College, London, UK), N. Golding, S. Hay (SEEG, Univ. Oxford, Oxford, UK) ad W. Wint (ERGO ltd, Oxford, UK), and S. Vanwambeke (UCL, Louvain-la-Neuve, Belgium).
Related publications
Point pattern simulation modelling of extensive and intensive chicken farming in Thailand: Accounting for clustering and landscape characteristics
C. Chaiban, C. Biscio, W. Thanapongtharm, M. Tildesley, X. Xiao, T. P. Robinson, S. O. Vanwambeke, and M. Gilbert.
"Agricultural Systems",
Vol. 173,
Pages 335-344,
2019.
Predictive gravity models of livestock mobility in Mauritania: The effects of supply, demand and cultural factors
G. Nicolas, A. Apolloni, C. Coste, G. R. W. Wint, R. Lancelot, and M. Gilbert.
"PLoS One",
Vol. 13,
Issue 7,
Pages e0199547,
2018.
Using random forest to improve the downscaling of global livestock census data
G. Nicolas, T. P. Robinson, G. R. W. Wint, G. Conchedda, G. Cinardi, and M. Gilbert.
"PLoS One",
Vol. 11,
Issue 3,
Pages e0150424,
2016.
Global mapping of highly pathogenic avian influenza H5N1 and H5Nx clade 2.3.4.4 viruses with spatial cross-validation
M. S. Dhingra, J. Artois, T. P. Robinson, C. Linard, C. Chaiban, I. Xenarios, R. Engler, R. Liechti, D. Kuznetsov, X. Xiao, S. V. Dobschuetz, F. Claes, S. H. Newman, G. Dauphin, and M. Gilbert.
"eLife",
Vol. 5,
Pages e19571,
2016.
Spatial characterization of colonies of the flying fox bat, a carrier of Nipah virus in Thailand
W. Thanapongtharm, C. Linard, W. Wiriyarat, P. Chinsorn, B. Kanchanasaka, X. Xiao, C. Biradar, R. G. Wallace, and M. Gilbert.
"BMC Veterinary Research",
Vol. 11,
Issue 1,
2015.
Income disparities and the global distribution of intensively farmed chicken and pigs
M. Gilbert, G. Conchedda, T. P. Van Boeckel, G. Cinardi, C. Linard, G. Nicolas, W. Thanapongtharm, L. D'Aietti, W. Wint, S. H. Newman, and T. P. Robinson.
"PLoS One",
Vol. 10,
Issue 7,
Pages e0133381,
2015.
Predicting the risk of avian influenza A H7N9 infection in live-poultry markets across Asia
M. Gilbert, N. Golding, H. Zhou, G. R. W. Wint, T. P. Robinson, A. J. Tatem, S. Lai, S. Zhou, H. Jiang, D. Guo, Z. Huang, J. P. Messina, X. Xiao, C. Linard, T. P. Van Boeckel, V. Martin, S. Bhatt, P. W. Gething, J. J. Farrar, S. I. Hay, and H. Yu.
"Nature Communications",
Vol. 5,
Pages 4116,
2014.